Warning
This document is for an in-development version of Galaxy. You can alternatively view this page in the latest release if it exists or view the top of the latest release's documentation.
September 2021 Galaxy Release (v 21.09)

Highlights
Tool Panel
We now have “Tool Panel Views”! These are different views into the same toolbox and might help it make it easier to find the tools you want. In the future, there are plans to create user-customisable toolboxes, but until then go explore the new EDAM Ontology-based toolboxes which organise tools around scientific areas or processes, or based on the scientific process they do. For example, a category like “Filtering” might have tools like “select lines” or “filter bam by quality”, both doing the same process of filtering, despite their different file types and formats.
Unfortunately not all tools are fully annotated yet, and while we also pull terms from bio.tools, this still doesn’t get us full coverage and many tools will still appear under a large section “Uncategorized”. Hopefully this will improve as users and tool developers annotate these tools with the appropriate terms that help everyone find them.
More details can be found in Pull Request 12327, Pull Request 12365, and Pull Request 12291
Collections (Beta History Updates)
Did you ever create a collection with the wrong dbkey or datatype? Well, in the Beta History you can now change collections’ datatypes and dbkeys! This should save a lot of time for everyone working with large datasets (Pull Request 11799).
Additionally, did a collection ever fail for you? And you wondered why? Now you can find out with the view details button for collections (Pull Request 12261)!
Importing Data
Selecting datasets from the Remote Files view has gotten easier! Now you can select folders and files, and import at once all of the datasets recursively in those folders. Previously you could only select files within a folder, so this is a huge improvement in the usability of such a key new feature of Galaxy (Pull Request 12310). But with great power comes great responsibility, be careful not to import the entirety of NCBI!
Reports
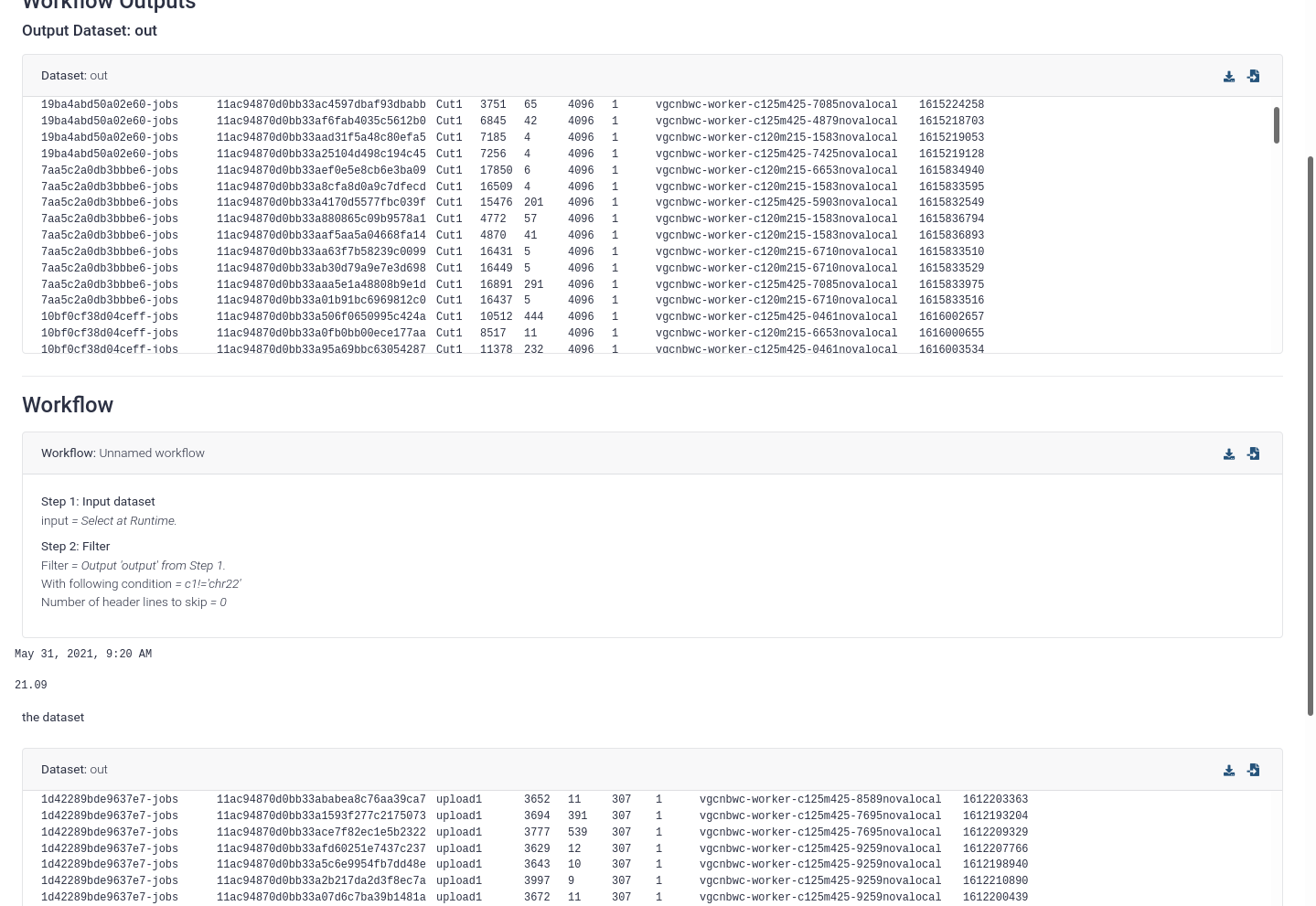
Report components used to automatically arrange themselves in a smart way, but due to some issues with the floating of report components these will now be full width until we can figure out a good way to provide a similar feature allowing you to control the layout of the report (Pull Request 12085).
News Webhook
You might be reading this in the recently added “News” feature! This is the very first iteration of this component, hopefully in the future we will expand it to bring you all of your news and Galaxy notifications. The idea with the current iteration is that whenever your server admin updates Galaxy to a new version, everyone using the server will get a small, non-intrusive notification with these release notes! There you can find out all of the new features in Galaxy and quick videos on their use. (Pull Request 12396)
New Datatypes
Added documentation for FASTQ datatypes and implemented quality check (thanks to @bernt-matthias). Pull Request 11931
Make dataset preview for h5mlm datatype (thanks to @qiagu). Pull Request 11935
Add datatypes for Structural Materials Hexrd application (thanks to @jj-umn). Pull Request 11957
Adding new subclass types (thanks to @maikenp). Pull Request 12097
Converters: use target datatype (thanks to @bernt-matthias). Pull Request 12185
Add bref3 datatype (thanks to @gallardoalba). Pull Request 12199
Converters: add missing tests and add linting to converter tests (thanks to @bernt-matthias). Pull Request 12202
Converters: Unify converters to tabix and bgzip (thanks to @bernt-matthias). Pull Request 12213
Converters: Unify molecules converters (thanks to @bernt-matthias). Pull Request 12214
Converters: Unify dcd, trr, xtc (thanks to @bernt-matthias). Pull Request 12224
converters: Unify bcf converters (thanks to @bernt-matthias). Pull Request 12225
Fix edta metadata setting (thanks to @bernt-matthias). Pull Request 12273
Increase specificity of mothur.pair.dist sniffer (thanks to @bernt-matthias). Pull Request 12280
Add “ExpressionSet RData object” Datatype (thanks to @mtekman). Pull Request 12336
Small fix in binary.py (thanks to @melibleq). Pull Request 12384
Parse sam metadata from sam files (thanks to @bernt-matthias). Pull Request 12392
Add ONNX datatype (thanks to @anuprulez). Pull Request 12429
Drop bcftools requirement from set_metadata tool Pull Request 12472
Fix cmap sniffer (thanks to @astrovsky01). Pull Request 12509
Add support for RDS files and improvements for RData (thanks to @bernt-matthias). Pull Request 12712
Add file_ext property to Pretext and others datatypes (thanks to @bernt-matthias). Pull Request 12713
Make vg a subclass of CompressedArchive (thanks to @bernt-matthias). Pull Request 12718
Yaml datatype backport Pull Request 12745
Builtin Tool Updates
Changes to Collection Operation Help sections and parameter names Pull Request 11068
GPU enabled jupyter notebook for machine learning powered by Jupyter lab and Tensorflow (thanks to @anuprulez). Pull Request 11484
Update bam.iobio interactive tool wrapper (thanks to @luke-c-sargent). Pull Request 11537
Add tool for exporting individual files to galaxy file source plugins. Pull Request 11613
NCBI Datasets data source tool Pull Request 11738
Fix typo in interactivetool_jupyter_notebook.xml help section (thanks to @maximskorik). Pull Request 12077
Update interactivetool_pyiron.xml (thanks to @gmauro). Pull Request 12127
Update AskOmics interactive tool to 4.3.1 (thanks to @abretaud). Pull Request 12159
Fix patterns in Grep1 tool Pull Request 12166
Remove unused legacy controller things Pull Request 12172
Add
<creator>to the tool schema template, use live links in xsd Pull Request 12242Systematic handling of remotely required tool files. Pull Request 12250
Restore Grep1 version 1.0.1 Pull Request 12252
Vuefy and improve granularity for tool HTML sanitization Pull Request 12283
Allow bio.tools mappings for legacy tools. Pull Request 12289
Allow skipping sanitization of / char in export_remote tool Pull Request 12372
Lock location file before adding values Pull Request 12446
Modernize sorter (sort1) tool Pull Request 12619
Improve error handling in DirectoryUriToolParameter validation (thanks to @davelopez). Pull Request 12760
Release Testing Team
A special thanks to the release testing team for testing many of the new features and reporting many bugs:
Release Notes
Please see the full release notes for more details.
To stay up to date with Galaxy’s progress, watch our screencasts; visit our community Hub; and follow us on Bluesky, Mastodon, and LinkedIn.
You can always chat with us on Matrix.
Thanks for using Galaxy!